SHAP with structured data classification
Explainable AI with TensorFlow, Keras and SHAP
This tutorial is mainly based on the Keras tutorial “Structured data classification from scratch” by François Chollet and “Census income classification with Keras” by Scott Lundberg.
- To keep this tutorial simple, we will only use numerical features in our binary classification example.
- Take a look at to see an example of how to use more advanced Keras data preprocessing layers.
Setup
import numpy as np
import pandas as pd
import tensorflow as tf
from tensorflow.keras import layers
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import StandardScaler
import shap
tf.__version__'2.8.1'# print the JS visualization code to the notebook
shap.initjs()Data
Here’s the description of each feature:
| Column | Description | Feature Type |
|---|---|---|
| Age | Age in years | Numerical |
| Sex | (1 = male; 0 = female) | Categorical |
| CP | Chest pain type (0, 1, 2, 3, 4) | Categorical |
| Trestbpd | Resting blood pressure (in mm Hg on admission) | Numerical |
| Chol | Serum cholesterol in mg/dl | Numerical |
| FBS | fasting blood sugar in 120 mg/dl (1 = true; 0 = false) | Categorical |
| RestECG | Resting electrocardiogram results (0, 1, 2) | Categorical |
| Thalach | Maximum heart rate achieved | Numerical |
| Exang | Exercise induced angina (1 = yes; 0 = no) | Categorical |
| Oldpeak | ST depression induced by exercise relative to rest | Numerical |
| Slope | Slope of the peak exercise ST segment | Numerical |
| CA | Number of major vessels (0-3) colored by fluoroscopy | Both numerical & categorical |
| Thal | normal; fixed defect; reversible defect | Categorical (string) |
| Target | Diagnosis of heart disease (1 = true; 0 = false) | Target |
We will only use the following continuous numerical features to predict the variable “Target” (diagnosis of heart disease):
agetrestbpscholthalacholdpeakslope
Data import
- Let’s download the data and load it into a Pandas dataframe:
file_url = "http://storage.googleapis.com/download.tensorflow.org/data/heart.csv"
df = pd.read_csv(file_url)Data preparation
# make target variable
y = df.pop('target')# prepare features
list_numerical = ['age', 'thalach', 'trestbps', 'chol', 'oldpeak']
X = df[list_numerical]Data splitting
- Let’s split the data into a training and test set
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state=42)Feature preprocessing
scaler = StandardScaler().fit(X_train[list_numerical])
X_train[list_numerical] = scaler.transform(X_train[list_numerical])
X_test[list_numerical] = scaler.transform(X_test[list_numerical])Model
Now we can build the model using the Keras sequential API:
model = tf.keras.Sequential([
tf.keras.layers.Dense(10, activation='relu'),
tf.keras.layers.Dense(10, activation='relu'),
tf.keras.layers.Dense(1, activation='sigmoid')
])model.compile(optimizer="adam",
loss ="binary_crossentropy",
metrics=["accuracy"])model.fit(X_train, y_train,
epochs=15,
batch_size=13,
validation_data=(X_test, y_test)
)Epoch 1/15
1/19 [>.............................] - ETA: 2s - loss: 0.6155 - accuracy: 0.9231
2022-06-12 17:00:50.797019: W tensorflow/core/platform/profile_utils/cpu_utils.cc:128] Failed to get CPU frequency: 0 Hz
19/19 [==============================] - 0s 4ms/step - loss: 0.6545 - accuracy: 0.7107 - val_loss: 0.6483 - val_accuracy: 0.7049
Epoch 2/15
19/19 [==============================] - 0s 1ms/step - loss: 0.6301 - accuracy: 0.7273 - val_loss: 0.6289 - val_accuracy: 0.7213
Epoch 3/15
19/19 [==============================] - 0s 1ms/step - loss: 0.6078 - accuracy: 0.7273 - val_loss: 0.6109 - val_accuracy: 0.7213
Epoch 4/15
19/19 [==============================] - 0s 1ms/step - loss: 0.5867 - accuracy: 0.7273 - val_loss: 0.5950 - val_accuracy: 0.7213
Epoch 5/15
19/19 [==============================] - 0s 1ms/step - loss: 0.5659 - accuracy: 0.7273 - val_loss: 0.5821 - val_accuracy: 0.7213
Epoch 6/15
19/19 [==============================] - 0s 1ms/step - loss: 0.5476 - accuracy: 0.7273 - val_loss: 0.5719 - val_accuracy: 0.7213
Epoch 7/15
19/19 [==============================] - 0s 1ms/step - loss: 0.5307 - accuracy: 0.7273 - val_loss: 0.5631 - val_accuracy: 0.7213
Epoch 8/15
19/19 [==============================] - 0s 1ms/step - loss: 0.5159 - accuracy: 0.7273 - val_loss: 0.5562 - val_accuracy: 0.7213
Epoch 9/15
19/19 [==============================] - 0s 1ms/step - loss: 0.5042 - accuracy: 0.7273 - val_loss: 0.5508 - val_accuracy: 0.7213
Epoch 10/15
19/19 [==============================] - 0s 1ms/step - loss: 0.4939 - accuracy: 0.7273 - val_loss: 0.5468 - val_accuracy: 0.7213
Epoch 11/15
19/19 [==============================] - 0s 1ms/step - loss: 0.4860 - accuracy: 0.7273 - val_loss: 0.5440 - val_accuracy: 0.7213
Epoch 12/15
19/19 [==============================] - 0s 951us/step - loss: 0.4794 - accuracy: 0.7273 - val_loss: 0.5422 - val_accuracy: 0.7213
Epoch 13/15
19/19 [==============================] - 0s 1ms/step - loss: 0.4737 - accuracy: 0.7273 - val_loss: 0.5412 - val_accuracy: 0.7213
Epoch 14/15
19/19 [==============================] - 0s 1ms/step - loss: 0.4685 - accuracy: 0.7273 - val_loss: 0.5400 - val_accuracy: 0.7213
Epoch 15/15
19/19 [==============================] - 0s 1ms/step - loss: 0.4649 - accuracy: 0.7314 - val_loss: 0.5386 - val_accuracy: 0.7213
<keras.callbacks.History at 0x283368a00>- Configure the model with Keras Model.compile:
Let’s visualize our connectivity graph:
# `rankdir='LR'` is to make the graph horizontal.
tf.keras.utils.plot_model(model, show_shapes=True, rankdir="LR")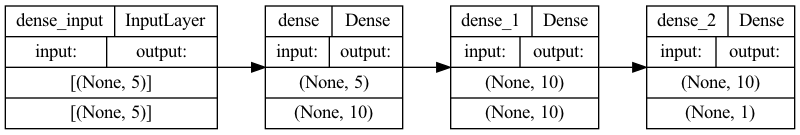
loss, accuracy = model.evaluate(X_test, y_test)
print("Accuracy", accuracy)2/2 [==============================] - 0s 1ms/step - loss: 0.5386 - accuracy: 0.7213
Accuracy 0.7213114500045776Perform inference
The model you have developed can now classify a row from a CSV file directly after you’ve included the preprocessing layers inside the model itself. Let’s demonstrate the process:
Save the heart diseases classification model
model.save('classifier_hd')INFO:tensorflow:Assets written to: classifier_hd/assets
2022-06-12 17:01:00.553972: W tensorflow/python/util/util.cc:368] Sets are not currently considered sequences, but this may change in the future, so consider avoiding using them.- Load model
reloaded_model = tf.keras.models.load_model('classifier_hd')predictions = reloaded_model.predict(X_train)print(
"This particular patient had a %.1f percent probability "
"of having a heart disease, as evaluated by our model." % (100 * predictions[0][0],)
)This particular patient had a 24.8 percent probability of having a heart disease, as evaluated by our model.SHAP
We use our model and a selection of 50 samples from the dataset to represent “typical” feature values (the so called background distribution).
explainer = shap.KernelExplainer(model, X_train.iloc[:50,:])Now we use 500 perterbation samples to estimate the SHAP values for a given prediction (at index location 20). Note that this requires 500 * 50 evaluations of the model.
shap_values = explainer.shap_values(X_train.iloc[20,:], nsamples=500)The so called force plot below shows how each feature contributes to push the model output from the base value (the average model output over the training dataset we passed) to the model output. Features pushing the prediction higher are shown in red, those pushing the prediction lower are in blue. To learn more about force plots, take a look at this Nature BME paper from Lundberg et al. (2018).
shap.force_plot(explainer.expected_value, shap_values[0], X_train.iloc[20,:])Visualization omitted, Javascript library not loaded!
Have you run initjs() in this notebook? If this notebook was from another user you must also trust this notebook (File -> Trust notebook). If you are viewing this notebook on github the Javascript has been stripped for security. If you are using JupyterLab this error is because a JupyterLab extension has not yet been written.
Explain many predictions
If we take many force plot explanations such as the one shown above, rotate them 90 degrees, and then stack them horizontally, we can see explanations for an entire dataset (see content below). Here, we repeat the above explanation process for 50 individuals.
To understand how a single feature effects the output of the model we can plot the SHAP value of that feature vs. the value of the feature for all the examples in a dataset. Since SHAP values represent a feature’s responsibility for a change in the model output, the plot below represents the change in the dependent variable. Vertical dispersion at a single value of represents interaction effects with other features.
shap_values50 = explainer.shap_values(X_train.iloc[50:100,:], nsamples=500) 0%| | 0/50 [00:00<?, ?it/s]shap.force_plot(explainer.expected_value, shap_values50[0], X_train.iloc[50:100,:])Visualization omitted, Javascript library not loaded!
Have you run initjs() in this notebook? If this notebook was from another user you must also trust this notebook (File -> Trust notebook). If you are viewing this notebook on github the Javascript has been stripped for security. If you are using JupyterLab this error is because a JupyterLab extension has not yet been written.